Poster Session 2026

- Home
- Amanda N. D. Adams
- Olivia Ambrose
- Prooksa Ananchuensook
- Victoria H Anderson
- Mariam Baig
- Suchandra Banerjee
- Ofri Bar
- Leah C Beauchamp
- Paige K Berger
- Chandrima Bhattacharya
- Katy Bond
- Camille Briskin
- Amanda Darling
- Mengxi Du
- Guilherme Fahur Bottino
- Elsa Fristot
- Emmanuel A Gyimah
- Erik Hasenoehrl
- Kyoo Heo
- Nathan T Jacobs
- Jordan S L Jensen
- Yehoon Jo
- Da Jung Jung
- Roka Kakehi
- Thomas M Kuntz
- S. Li
- Valeria Lugo Mesa
- Xochitl C Morgan
- Jacob T Nearing
- Ana Nogal
- Maribel Okiye
- Wakako Okuda
- Lily A Palumbo
- Yiming Shi
- Jack T Sumner
- Vishnu Thayil Valappil
- Chahat Upreti
- Maggie Viland
- Dongyu Wang
- Ya Wang
- Xinyu Wang
- Yan Yan
- Yiyan Yang
Poster Session 2026
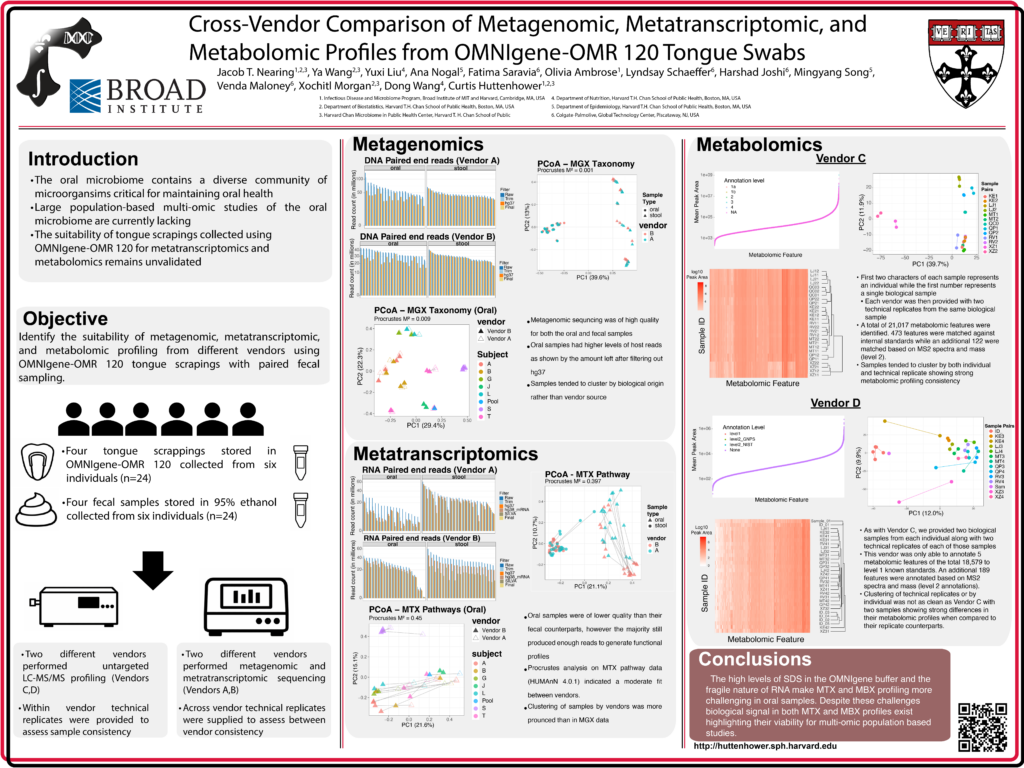
Cross‑Vendor Comparison of Metagenomic, Metatranscriptomic, and Metabolomic Profiles from OMNIgene‑OMR 120 Tongue Swabs
Presented By: Jacob T. Nearing
The oral microbiome contains a diverse community of microorganisms that play critical roles in maintaining oral health. However, large population‑based oral microbiome efforts integrating multiomics data remain limited. Most prior studies have relied on 16S rRNA profiling, have focused on the saliva microbiome, and/or have lacked information on microbial transcriptional activity and small‑molecule chemistry. Further, the suitability of tongue swabs, which contain high human DNA content, for metatranscriptomic (MTX) sequencing and metabolomic (MBX) profiling remains poorly validated.
To identify the suitability for MTX and MBX on tongue swabs, we conducted a pilot study using two sets of 24 paired tongue and fecal samples collected from six individuals, preserved in OMNIgene‑OMR 120 oral sampler (tongue) or 95% ethanol (feces). One set of samples was sent to two different vendors (vendors A & B) for paired MGX and MTX sequencing and a second set was sent to two different vendors (vendors C& D) for untargeted LC-MS/MS analysis.
Oral and fecal sequencing data were generally high quality from both vendors (MGX > 20 million reads, MTX > 10 million reads after data quality control), although a small number of tongue swabs samples failed to generate MGX (vendor A: n=3) or MTX (vendor A: n=5; vendor B: n=2) data due to either low read count or failed extraction. Microbial taxonomic profiles from both fecal and oral samples clustered primarily by sample and subject rather than vendor (fecal Procrustes M²=0.006; oral M²=0.009), with similar results at the functional pathway level (fecal M²=0.002; oral M²=0.016). In contrast, MTX data, particularly from oral samples, showed higher variability attributable to vendor choice (oral M²=0.45). Fecal MBX profiles showed strong technical replicate clustering and agreement with previously published fecal MBX data. Oral MBX was less optimized: vendor C confidently matched only 5 of 18,579 features to internal standards, with many lower‑confidence features likely originating from the OMNIgene buffer rather than the biological sample. Vendor D showed strong replicate clustering and matched 473 of 21,019 features to internal standards. Moreover, several oral relevant metabolites received high levels of annotation confidence through either internal standard matching or MS2 spectra matching. Overall, our findings indicate that although oral samples pose greater challenges for MTX and MBX analyses, they remain viable for multi‑omics characterization in population‑based studies.